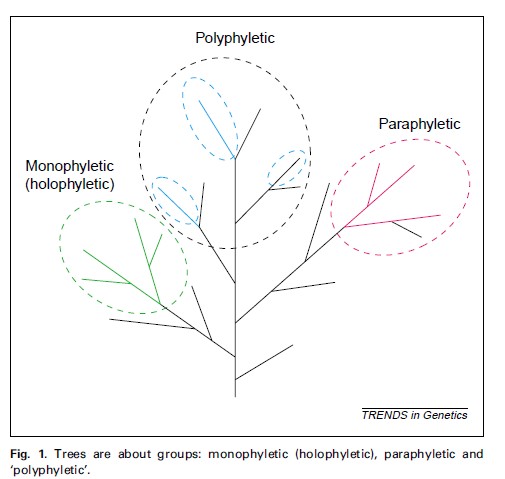
进化树的应用越来越广,很多不同研究背景的人都非常了解进化树到底能告诉我们哪些信息。现在网上也有各种各样的文章和博客介绍如何构建进化树。PLoB中也有一些文章专门介绍如何构建进化树,例如:
以上都是教大家构建进化树软件的使用方法,但是如何去选择数据?构建进化树的方法也有很多种,不同的数据如何去选择算法?构建进化树之后如何去分析构建好的进化树呢?
下面介绍一篇关于Phylogeny入门级的文章,这篇入门论文的目的是要介绍一些基本的建立系统进化树的方法,特别是为那些正在看别人建立的树或是正在开始学习建立树的人。该文内容包括了:怎样读懂一个树,怎样获得数据,多个序列的比对,系统进化的方法,自检分析和分枝长度等问题,以及相关的软件和资源。
Phylogeny for the faint of heart: a tutorial
Sandra L. Baldauf
Department of Biology, University of York, Box 373, York, UK YO10 5YW
Abstract
Phylogenetic trees seem to be finding ever broader applications, and researchers from very different backgrounds are becoming interested in what they might have to say. This tutorial aims to introduce the basics of building and interpreting phylogenetic trees. It is intended for those wanting to understand better what they are looking at when they look at someone else's trees or to begin learning how to build their own. Topics covered include: how to read a tree, assembling a dataset, multiple sequence alignment (how it works and when it does not), phylogenetic methods, bootstrap analysis and long-branch artefacts, and software and resources.
Phylogenetics is the science of estimating the evolutionary past, in the case of molecular phylogeny, based on the comparison of DNA or protein sequences. The idea of representing these hypotheses as trees probably dates back to Darwin, but the numerical calculation of trees using quantitative methods is relatively recent, and their application to molecular data even more so. In the age of rapid and rampant gene sequencing, molecular phylogeny has truly come into its own, emerging as a major tool for making sense of a sometimes overwhelming amount information.
This tutorial aims to introduce the basic principles behind and programs for constructing evolutionary trees (phylogenetic analysis). It is intended primarily for those who want to read other people's trees, but also as a general introduction for those who might wish to begin to try building their own. In the latter case the reader is warned – phylogenetic analysis and evolutionary theory are not trivial pursuits; as with any new methodology, it is advisable to seek expert help before getting in too deep.
全文链接:http://www.sciencedirect.com/science/article/pii/S0168952503001124