在之前的一节当中,图型名称有些混乱,从这一节开始将做如下统一(不全面):
| 英文名称 | 中文名称 |
| bar | 条形图 |
| line | 线图 |
| area | 面积图 |
| pie | 饼图 |
| high-low | 高低图 |
| pareto | 帕累托图 |
| control | 控制图 |
| boxplot | 箱线图 |
| error bar | 误差条图 |
| scatter | 散点图 |
| P-P | P-P正态概率图 |
| Q-Q | Q-Q正态概率图 |
| sequence | 序列图 |
| ROC Curve | ROC分类效果曲线图 |
| Time Series | 时间序列图 |
好了,言归正传。那么什么又是点柱图(dot histogram)呢?之前我又称之为蜂群图(beeswarm)。还有称之为抖点图(jitter plots)。总之无论如何,在糗世界里我都称之为点柱图吧。
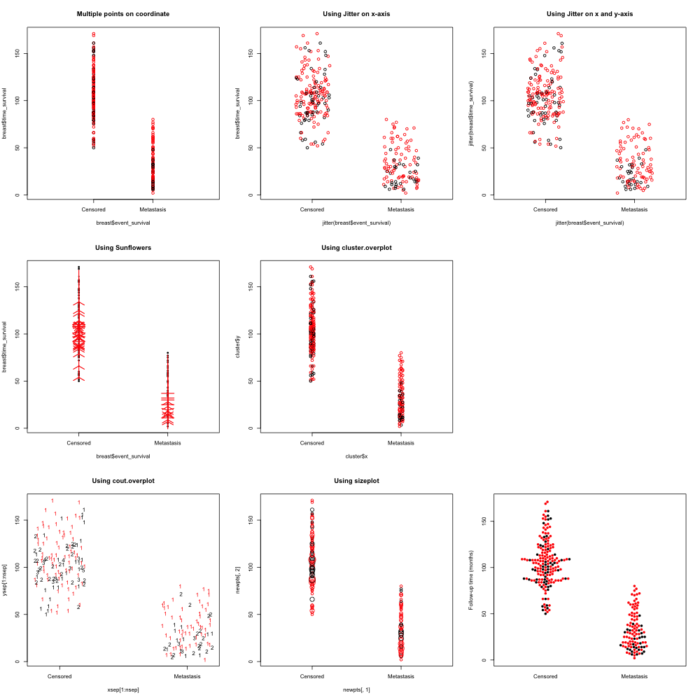
我们先看点柱图效果:
以下是代码
> require(beeswarm)
> data(breast)
> head(breast)
ER ESR1 ERBB2 time_survival event_survival
100.CEL.gz neg 8.372133 13.085894 39 1
103.CEL.gz pos 10.559356 9.491683 97 0
104.CEL.gz pos 12.299905 9.599574 11 1
105.CEL.gz pos 10.776632 9.681747 99 0
106.CEL.gz pos 10.505124 9.436763 40 1
107.CEL.gz neg 10.377741 8.695576 94 0
> require(plotrix)
> cluster<-cluster.overplot(breast$event_survival, breast$time_survival)
> png("dothist.png",width=1000,height=1000)
> opar<-par(mfrow=c(3,3))
> plot (breast$event_survival, breast$time_survival, main="Multiple points on coordinate",col=as.numeric(breast$ER),xaxt="n",xlim=c(-1,2))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> plot(jitter(breast$event_survival), breast$time_survival, main="Using Jitter on x-axis",col=as.numeric(breast$ER),xaxt="n",xlim=c(-0.5,1.5))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> plot(jitter(breast$event_survival), jitter(breast$time_survival), main="Using Jitter on x and y-axis",col=as.numeric(breast$ER),xaxt="n",xlim=c(-0.5,1.5))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> sunflowerplot(breast$event_survival, breast$time_survival, main="Using Sunflowers",xaxt="n",xlim=c(-0.5,1.5))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> plot(cluster, main="Using cluster.overplot",col=as.numeric(breast$ER),xaxt="n",xlim=c(-0.5,1.5))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> count.overplot(jitter(breast$event_survival), jitter(breast$time_survival), main="Using cout.overplot",col=as.numeric(breast$ER),xaxt="n")
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> sizeplot(breast$event_survival, breast$time_survival, main="Using sizeplot",col=as.numeric(breast$ER),xaxt="n",xlim=c(-0.5,1.5))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> beeswarm(time_survival ~ event_survival, data = breast,
+ method = 'swarm',
+ pch = 16, pwcol = as.numeric(ER),
+ xlab = '', ylab = 'Follow-up time (months)',
+ labels = c('Censored', 'Metastasis'))
> dev.off()
quartz
2
> par(opar) |
以下是解释
在很多情况下,我们画散点图的时候,有许多点拥有相同的横坐标,如果我们简单的使用plot(x,y)的方式,会显得这些点拥挤在一起,象图中左上角一样,非常的不舒服。我们需要把这些点分散开。
最基本的思路是,把横坐标抖散(jitter),使本来都拥有相同坐标的点的横坐标稍有不同。jitter是基类函数{base},无需调用任何包。
> plot(jitter(breast$event_survival), breast$time_survival, main="Using Jitter on x-axis",col=as.numeric(breast$ER),xaxt="n",xlim=c(-0.5,1.5))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis"))
> plot(jitter(breast$event_survival), jitter(breast$time_survival), main="Using Jitter on x and y-axis",col=as.numeric(breast$ER),xaxt="n",xlim=c(-0.5,1.5))
> axis(1,at=c(0,1),labels=c("Censored","Metastasis")) |
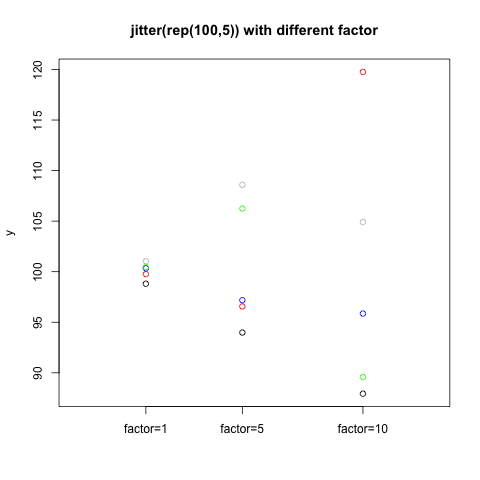
我们比较图中上边靠右的两个图,我们发现,如果只抖散x坐标的话,还是有些点会粘在一起,所以同时抖散y坐标会好一些。我们可以使用factor参数来控制jitter抖散的强度。
> plot(rep(c(1,5,10),each=5), c(jitter(rep(100,5),factor=1), jitter(rep(100,5),factor=5), jitter(rep(100,5), factor=10)), col=c("red","blue","green","gray","black"), xlim=c(-2,13), xlab="", ylab="y", xaxt="n", main="jitter(rep(100,5)) with different factor")
> axis(1,at=c(1,5,10),labels=c(paste("factor=",c(1,5,10),sep=""))) |
在graphics包中提供了一个sunflowerplot的函数。它的目的是用花瓣数目多少来显示落在同一坐标上的点的数目。但是从中左图看来,点多的时候效果并非总是那么好。
在plotrix包中提供了一些有意思的函数来解决点挤在一起的这个问题,它们分别是cluster.overplot, count.overplot, sizeplot。这三个函数的效果如图中及下靠左的两个。cluster.overplot的方法类似抖散,count overplot的方法是使用数字来显示落在同一坐标上的点的数目,sizeplot的方法是使用不同大小是点来显示落在同一坐标上的点的数目。从效果来看,点多的时候效果也并非理想。
而上一次提到过的蜂群图似乎是解决这一问题的最佳方案。
我们得出结论,在点数不同的情况下,使用plotrix包及sunflowerplot是不错的。但点数较多的情况下,还是使用jitter和beeswarm较为稳妥。
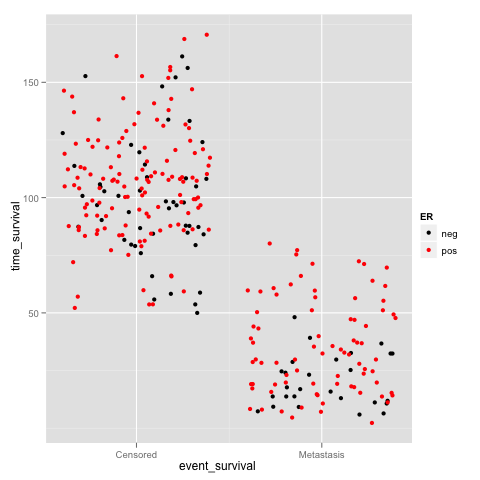
我们也可以使用ggplot2包中的geom来绘制点柱图
> require(beeswarm)
> data(breast)
> library(ggplot2)
> p<-ggplot(breast, aes(event_survival,time_survival))
> print(p+geom_jitter(aes(color=ER))+scale_colour_manual(value = c("black", "red")) + scale_x_continuous(breaks = c(0:1), labels = c("Censored", "Metastasis"))) |
还有一种图,名称为Engelmann-Hecker-Plot, 由plotrix的ehplot来实现。
> data(iris);library(plotrix) > ehplot(iris$Sepal.Length, iris$Species, + intervals=20, cex=1.8, pch=20, main="pch=20") > ehplot(iris$Sepal.Width, iris$Species, + intervals=20, box=TRUE, median=FALSE, main="box=TRUE") > ehplot(iris$Petal.Length, iris$Species, + pch=17, col="red", log=TRUE, main="pch=17") > ehplot(iris$Petal.Length, iris$Species, + offset=0.06, pch=as.numeric(iris$Species), main="pch=as.numeric(iris$Species)") > rnd <- sample(150) > plen <- iris$Petal.Length[rnd] > pwid <- abs(rnorm(150, 1.2)) > spec <- iris$Species[rnd] > ehplot(plen, spec, pch=19, cex=pwid, + col=rainbow(3, alpha=0.6)[as.numeric(spec)], main="cex and col changes") |