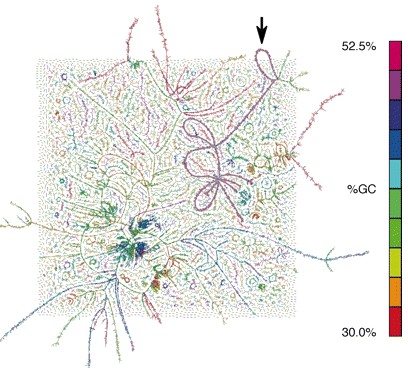
从由细菌、古细菌和病毒组装的错综复杂宏基因组DNA序列中,研究人员获得一种神秘的广古菌的整个基因组序列。
宏基因组学(metagenomics,译者注:也常译作环境基因组学)就是取样---一勺泥土或一杯海水---然后对样品内所有DNA进行测序。这种技术有助于研究人员评估在给定环境发现的生物多样性,但是直到现在它不能有效地对单个物种进行测序,因为没有办法对测得的数据进行解析而区别开每种物种基因组DNA序列。
但是2012年2月2日发表在《科学》期刊上的一种新计算方法可能提供答案和允许研究人员从宏基因组数据中对单个物种基因组测序。研究人员将他们的方法应用到从普吉湾(Puget Sound)水面获得的水样品上,能够从14种不同生物组成的DNA混合物中解析出两个完整的基因组序列。
美国华盛顿大学Virginia Armbrust也是该研究论文的共同作者。她对《自然》期刊说,“我们的方法令人激动,因为它最终为整个微生物群落如何工作打开一扇窗户。”这种技术也将允许研究人员对至今为止还不能在实验室培养条件下生长的生物进行测序。确实,Armbrust和她的同事们组装的一个基因组序列就是来自一种广古菌(Euryarchaeota)物种的,这种海洋微生物人们从未能够在实验室中培养。
Untangling Genomes from Metagenomes: Revealing an Uncultured Class of Marine Euryarchaeota
Vaughn Iverson, Robert M. Morris, Christian D. Frazar, Chris T. Berthiaume, Rhonda L. Morales, E. Virginia Armbrust
Ecosystems are shaped by complex communities of mostly unculturable microbes. Metagenomes provide a fragmented view of such communities, but the ecosystem functions of major groups of organisms remain mysterious. To better characterize members of these communities, we developed methods to reconstruct genomes directly from mate-paired short-read metagenomes. We closed a genome representing the as-yet uncultured marine group II Euryarchaeota, assembled de novo from 1.7% of a metagenome sequenced from surface seawater. The genome describes a motile, photo-heterotrophic cell focused on degradation of protein and lipids and clarifies the origin of proteorhodopsin. It also demonstrates that high-coverage mate-paired sequence can overcome assembly difficulties caused by interstrain variation in complex microbial communities, enabling inference of ecosystem functions for uncultured members.