Small non-coding RNAs近年来一直研究比较热门,尤其是在真核生物中。因此PLoB网站上面也专门开辟了ncRNA专栏来介绍这一块的研究进展。最近在做一个细菌的转录组课题,需要鉴定其中的sRNA。我的鉴定思路是从三个角度同时分析:1)从已有的论文中收集;2)基于现有的数据库搜索;3)De novo 预测
今天主要给大家推荐一个比较不错的细菌Small non-coding RNAs数据库,sRNAMAP。这个数据库中收集了不少细菌中的ncRNA和调节性RNA,更重要的一点是该数据库包含了其中一些RNA作用的target。同时该数据库还能为为用户的序列在线注释non-coding RNA的功能,不过我是直接把所有的ncRNA序列下载下来自己用BLAT程序比对的。有需要的童鞋可以留意一下这个数据库。
下面是他们的数据库的网站和相关论文即摘要:
网址地址:http://srnamap.mbc.nctu.edu.tw/
论文:
摘要:
sRNAMap: genomic maps for small non-coding RNAs, their regulators and their targets in microbial genomes.
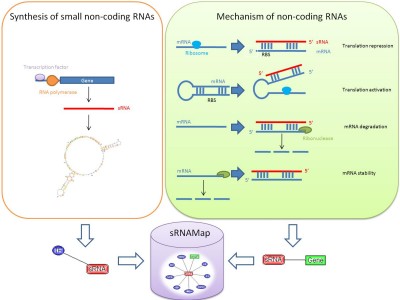
Small non-coding RNAs (sRNAs) carry out a variety of biological functions and affect protein synthesis and protein activities in prokaryotes. Recently, numerous sRNAs and their targets were identified in Escherichia coli and in other bacteria. It is crucial to have a comprehensive resource concerning the annotation of small non-coding RNAs in microbial genomes. This work presents an integrated database, namely sRNAMap, to collect the sRNA genes, the transcriptional regulators of sRNAs and the sRNA target genes by integrating a variety of biological databases and by surveying literature. In this resource, we collected 397 sRNAs, 62 regulators/sRNAs and 60 sRNAs/targets in 70 microbial genomes. Additionally, more valuable information of the sRNAs, such as the secondary structure of sRNAs, the expressed conditions of sRNAs, the expression profiles of sRNAs, the transcriptional start sites of sRNAs and the cross-links to other biological databases, are provided for further investigation. Besides, various textual and graphical interfaces were designed and implemented to facilitate the data access in sRNAMap. sRNAMap is available at http://sRNAMap.mbc.nctu.edu.tw/.