一、安装
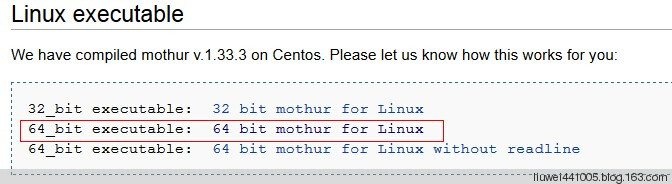
下载 http://www.mothur.org/wiki/Download_mothur, 我下载的是中间那个,但不知道最后一个的readline是干嘛的,有高手可以指点下。
下载后得到压缩文件Mothur_cen_64.zip,上传到服务器,解压:unzip Mothur_cen_64.zip,得到文件夹mothur,为了方便使用,可以将该文件夹的路径添加到环境变量中(编辑.bashrc文件)。
二、运行方式
1、Interactive mode 交互式
在terminal输入mothur进入交互式界面,
mothur > mothur > align.seqs(help) mothur > quit
2、Batch mode 批量运行模式
http://www.mothur.org/wiki/Batch_mode
把命令写到一个文件中batchfile
--------------------------------batchfile-------------------------------------- cluster(phylip=98_sq_phylip_amazon.dist, cutoff=0.1) collect.single() rarefaction.single() --------------------------------------------------------------------------------
运行:
../mothur/mothur batchfile
3、Command line mode 命令行模式
http://www.mothur.org/wiki/Command_line_mode
mothur "#read.dist(phylip=98_sq_phylip_amazon.dist, cutoff=0.1); cluster(); collect.single()"
命令间用分号分隔,所有的命令用双引号括起来,且括号内的命令以#号开头
三、Analysis examples
http://www.mothur.org/wiki/Analysis_examples
1、MiSeq SOP (standard operating procedure)http://www.mothur.org/wiki/MiSeq_SOP
(1)paire end根据overlap合并
首先做一个文件包含如下内容
--------------------------------stability.files----------------------------- F3D0 F3D0_S188_L001_R1_001.fastq F3D0_S188_L001_R2_001.fastq F3D141 F3D141_S207_L001_R1_001.fastq F3D141_S207_L001_R2_001.fastq F3D142 F3D142_S208_L001_R1_001.fastq F3D142_S208_L001_R2_001.fastq F3D143 F3D143_S209_L001_R1_001.fastq F3D143_S209_L001_R2_001.fastq F3D144 F3D144_S210_L001_R1_001.fastq F3D144_S210_L001_R2_001.fastq F3D145 F3D145_S211_L001_R1_001.fastq F3D145_S211_L001_R2_001.fastq F3D146 F3D146_S212_L001_R1_001.fastq F3D146_S212_L001_R2_001.fastq F3D147 F3D147_S213_L001_R1_001.fastq F3D147_S213_L001_R2_001.fastq F3D148 F3D148_S214_L001_R1_001.fastq F3D148_S214_L001_R2_001.fastq F3D149 F3D149_S215_L001_R1_001.fastq F3D149_S215_L001_R2_001.fastq F3D150 F3D150_S216_L001_R1_001.fastq F3D150_S216_L001_R2_001.fastq F3D1 F3D1_S189_L001_R1_001.fastq F3D1_S189_L001_R2_001.fastq F3D2 F3D2_S190_L001_R1_001.fastq F3D2_S190_L001_R2_001.fastq F3D3 F3D3_S191_L001_R1_001.fastq F3D3_S191_L001_R2_001.fastq F3D5 F3D5_S193_L001_R1_001.fastq F3D5_S193_L001_R2_001.fastq F3D6 F3D6_S194_L001_R1_001.fastq F3D6_S194_L001_R2_001.fastq F3D7 F3D7_S195_L001_R1_001.fastq F3D7_S195_L001_R2_001.fastq F3D8 F3D8_S196_L001_R1_001.fastq F3D8_S196_L001_R2_001.fastq F3D9 F3D9_S197_L001_R1_001.fastq F3D9_S197_L001_R2_001.fastq Mock Mock_S280_L001_R1_001.fastq Mock_S280_L001_R2_001.fastq -----------------------------------------------------------------------------
mothur "#make.contigs(file=stability.files, processors=4)"
程序运行时需要调用libreadline.so.6,#很重要的,不要遗漏。该程序运行时需要调用libreadline.so.6动态链接库。
查看merge后reads的统计信息:
summary.seqs(fasta=stability.trim.contigs.fasta)
对merge后的reads进行过滤:
screen.seqs(fasta=stability.trim.contigs.fasta, group=stability.contigs.groups, maxambig=0, maxlength=275)
or
screen.seqs(fasta=stability.trim.contigs.fasta, group=stability.contigs.groups, summary=stability.trim.contigs.summary, maxambig=0, maxlength=275) #Faster than upper cmd
查看当前的有效对象:
get.current() unique.seqs(fasta=stability.trim.contigs.good.fasta)
原文来自:http://liuwei441005.blog.163.com/blog/static/135705811201453094227246
更多关于Mothur使用介绍:



1F
Readline提供编译时的include与lib, 如果计算机上已经安装有readline,可以选without readline的那个源码包。如果没有安装或者编译mothur需要某个版本的readline,那就选with readline的源码包
2F
请教一下,数据库文件放在哪里可保证被mothur随时调用?